SARS-CoV-2 virus sequencing is tedious and labour-intensive, says specialist
 Credit: Adobe Stock
Credit: Adobe Stock
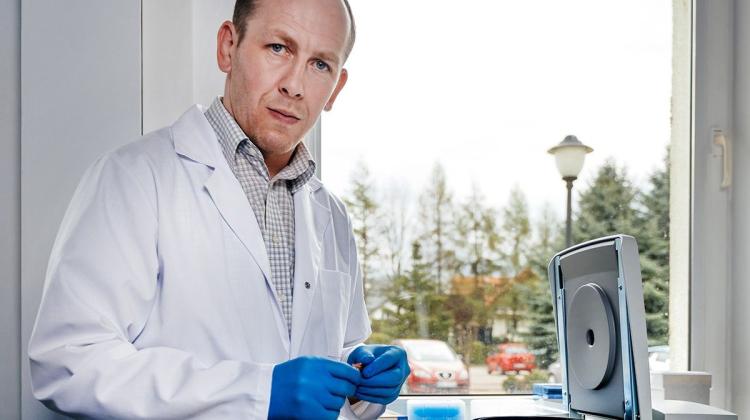
Determining the genetic structure of the SARS-CoV-2 virus and its mutations is a labour-intensive and tedious process, says Sylwia Krzętowska, head of the Epidemiological and Clinical Research Laboratory in Olsztyn. The nanopore sequencer (a device for sequencing nucleic acids) in Olsztyn analyses samples from four provinces.
The Olsztym lab processes samples from the following provinces: Warmia-Mazury, Kujawy-Pomorze, Pomorze and Podlasie.
Sylwia Krzętowska explains that the virus sequencing process begins with the delivery of a sample with the patient's material (after the PCR test) from a COVID lab. Not every sample with a positive result is suitable for sequencing. The basic criterion of sample usefulness is a sufficient amount of the virus RNA.
She said: “Sequencing is a study that has its own procedure. You need to perform specific actions to isolate the genetic material of the virus. They take about 2.5 days.
“First, you need to purify the sample so that only the SARS-CoV-2 genetic material remains. The human throat is not a clean place. A swab contains human epithelial cells, our DNA and DNA and RNA of microorganisms found in the respiratory tract. Purification of the sample is therefore a multi-stage, long, manual process.”
She added that the purified sample is then subjected to sequencing reactions in a sequencer, saying: “In the Olsztyn laboratory, we conduct Oxford nanopore sequencing. This method consists in the application of a film with nanopores on a special plate. Divided fragments of the genetic material of the SARS-CoV-2 virus pass through nanopores and thanks to the occurring electrochemical processes we can read the nucleotides of which the virus genome is built.”
Sequencing in the device takes about 72 hours. This time can be shortened to 48 hours, provided there is a sufficient amount of read genome sequences for the tested virus.
Once a specific nucleotide string is obtained for a given sample, the sequence is added to the international GISAID database, which verifies its similarity to the starting variant of the virus from Wuhan. Then, the virus variant and the type of mutation is finally determined. It can be, for example, the alpha (British), beta (South African), gamma (Brazilian) and delta (Indian) variant.
Krzętowska said: “Each fragment of the virus RNA is read about 20 times. The point is to obtain a sufficient number of sequence repetitions. Therefore, large computing power is needed.”
The feature of coronaviruses is that they mutate all the time. Most changes are harmless, while some cause specific mutations that are dangerous for people.
Dangerous viruses are characterized by greater infection rate, they may affect the severe course of the disease. Dangerous mutations also include those that cause the virus to become resistant to vaccination and previously approved medicines.
When asked if such a long sequencing process will not cause the laboratory to be overloaded in the case of the fourth wave of the pandemic, Krzętowska said that there is no need to analyse all samples.
She said: “We currently analyse all samples because there are few infections. According to the WHO and ECDC recommendations, we should analyse 5 to 10 percent of all SARS-CoV-2 positive samples. If we know that the outbreak occurs in a family that lives together, then there is no need to analyse samples from all family members. Similarly in the event of an outbreak in a workplace. There is a large probability that in these outbreaks the variant of the virus will be the same.”
It was similar with the British variant, which at some point began to dominate over other variants and, depending on the province, accounted for up to 97-98 percent of all occurring SARS-CoV-2 virus variants.
PAP – Nauka w Polsce, Agnieszka Libudzka
ali/ agt/ kap/
tr. RL
Przed dodaniem komentarza prosimy o zapoznanie z Regulaminem forum serwisu Nauka w Polsce.